human skin fibroblast cell line ws1 (ATCC)
Structured Review
Human Skin Fibroblast Cell Line Ws1, supplied by ATCC, used in various techniques. Bioz Stars score: 96/100, based on 391 PubMed citations. ZERO BIAS - scores, article reviews, protocol conditions and more
https://www.bioz.com/result/human skin fibroblast cell line ws1/product/ATCC
Average 96 stars, based on 391 article reviews
Images
1) Product Images from "Therapeutic Eradication of Cancer-associated Fibroblasts Inhibits in vivo progression of Pancreatic Cancer"
Article Title: Therapeutic Eradication of Cancer-associated Fibroblasts Inhibits in vivo progression of Pancreatic Cancer
Journal: bioRxiv
doi: 10.1101/2025.11.04.686484

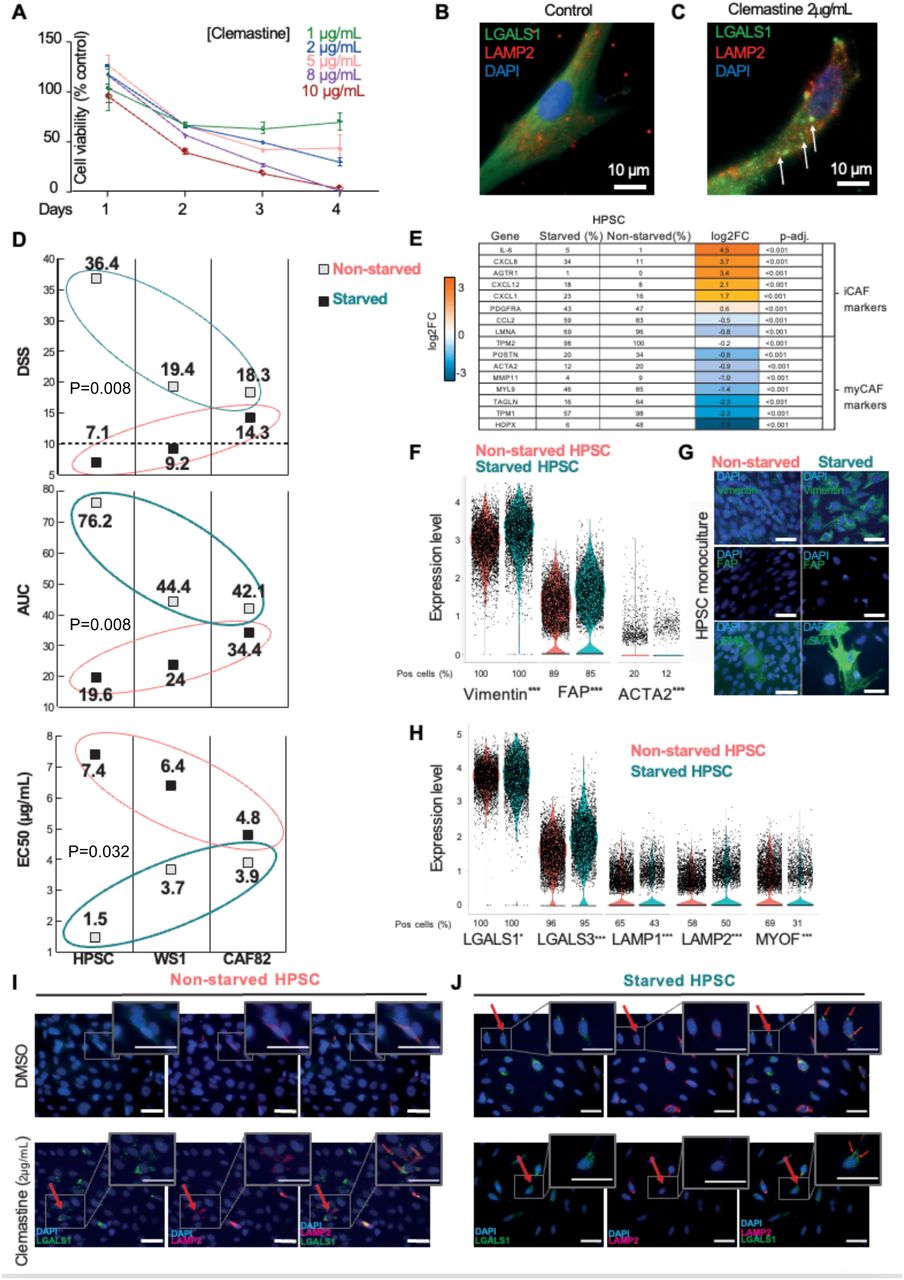
Figure Legend Snippet: A. WS-1 human fibroblast viability for indicated clemastine concentrations and timepoints. B-C. Representative Immunofluorescence micrographs of control untreated (B, N=10) and clemastine treated (2 µg/mL, 18 h, N=10) (C) WS-1 human fibroblasts. Lysosomal LAMP2 (red) and LGALS1 (green) are visualized. LAMP2 + /LGALS1 + permeable lysosomes are identified by arrows. Cell nuclei counterstained with DAPI (blue). Scalebar: 10 µm. D. Clemastine DSS, AUC and EC50 scores in three fibroblast lines (hPSC, WS1, CAF82) in their starved and non-starved states. E. Table of putative myCAF/iCAF marker genes and the percentages of cells positive for these in the starved and non-starved HPSC cultures. Average (avg) log2 fold change (FC) for the transcription level differences in activated iCAF-like cells versus parental HPSCs. Adjusted p-values. The change in transcription level (avg log2FC) ranges from blue (downregulated) to orange (upregulated). F. Violin plots of scRNAseq gene expression levels of CAF markers vimentin, FAP, and ACTA2 (gene coding for the α-SMA protein) in parental, non-starved (red) and starved, iCAF-like transformed, hPSCs (turquoise). G. Representative micrographs of FAP, vimentin, and α-SMA in non-starved (red) and starved (turquoise) HPSCs visualized by IF staining. H . Violin plots of scRNAseq gene expression levels of lysosomal markers in non-starved (red) and starved (turquoise) HPSCs. I-J . IF staining of LGALS1 (green) and LAMP2 (red) in untreated (DMSO) and clemastine-treated non-starved (I) and starved (J) HPSCs. LAMP2 + /LGALS1 + permeable lysosomes are identified by arrows. Clemastine treatment causes cytoplasmic LGALS1 to relocate to damaged lysosomes. Clemastine treatment also upregulated LAMP2 in HPSCs, but not in iCAF-like cells. LGALS1 relocation is also observed in untreated iCAF-like cells. *p<0.05, **p<0.001, ***p<0.0001.
Techniques Used: Immunofluorescence, Control, Marker, Gene Expression, Transformation Assay, Staining





![VPS41 negatively regulates KAI1 expression through the lysosomal pathway to confer resistance to apoptosis. A) Flowchart for screening VPS41 interaction proteins via differential protein analysis and mass spectrometry after VPS41 upregulation post‐irradiation. B) Volcano plot showing proteomic analysis (VPS41 vs Vector). C) Electron microscopy analysis reveals inhibited apoptosis progression in cells with upregulated VPS41 after irradiation (scale bar: 5 µm). D) Intersection of differential proteins identified four candidates: ISG15, KAI1, IFT20, and ATPAF1. E) Co‐localization of VPS41‐EGFP and KAI1‐BFP plasmids in WS1 and HaCaT cells assessed by confocal microscopy (scale bar: 20 µm). F) Immunoprecipitation confirms VPS41 binds KAI1. G) PNGase F treatment has minimal effect on VPS41‐KAI1 interaction. H) Western Blot shows upregulation of VPS41 decreases KAI1 expression, suppressing apoptosis, while VPS41 downregulation increases KAI1 expression and enhances apoptosis. I) VPS41 and KAI1 expression changes in HaCaT cells treated with CQ (20µ m ) or MG‐132 (20 µ m ) combined with X‐ray (10 Gy). J) Analysis of KAI1 decay rate after CHX (300 µ m ) treatment and X‐ray (10 Gy) in HaCaT cells. K) Effect of VPS41 knockdown and eccDNA transfection on apoptosis rates in irradiated cells with or without KAI1 overexpression. L) Apoptosis testing shows KAI1 reverses VPS41‐mediated radiation resistance. M) LDH measurement assesses the role of KAI1 in reversing VPS41‐mediated radiation resistance. P values were calculated using different statistical methods based on data type: unpaired two‐tailed t test for differential protein analysis and one‐way ANOVA followed by Bonferroni's post hoc test for multi‐group comparisons. Statistically significant differences are denoted as follows: * p < 0.05, ** p < 0.01. Data are presented as mean ± SD ( n = 3) unless otherwise specified. [Correction added on 28 April 2025, after first online publication: figure 5 is updated in this version].](https://pub-med-central-images-cdn.bioz.com/pub_med_central_ids_ending_with_4964/pmc12224964/pmc12224964__ADVS-12-2501934-g007.jpg)
